This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison .
Post-translational modifications____________________________________________________________________
After mRNA is translated by ribosomes in the cell to form a protein, the said protein may still be modified, which leads to even more functional diversity in proteins than what is already present at the genomic level [1]. Examples of post-translational modifications include phosphorylation, glycosylation, ubiquitination, SUMOylation, methylation, etc.. These modifications may regulate cellular activity through processes like protein degradation, transcriptional regulation and changing protein conformations.
After mRNA is translated by ribosomes in the cell to form a protein, the said protein may still be modified, which leads to even more functional diversity in proteins than what is already present at the genomic level [1]. Examples of post-translational modifications include phosphorylation, glycosylation, ubiquitination, SUMOylation, methylation, etc.. These modifications may regulate cellular activity through processes like protein degradation, transcriptional regulation and changing protein conformations.
Post-translational modification in DRD2______________________________________________________________________
There are various tools available online that can be used to predict post-translational modification in proteins based on their amino acid sequences. ExPasy—a bioinformatics resource portal—provides a list of the said tools.
There are various tools available online that can be used to predict post-translational modification in proteins based on their amino acid sequences. ExPasy—a bioinformatics resource portal—provides a list of the said tools.
|
NetNGlyc can be used to compare predicted N-glycosylation sites across different species and in different DRD2 protein isoforms. Glycosylation is a post-translational modification reaction whereby a carbohydrate (sugar) molecule is attached to proteins. In N-glycosylation—a process that occurs in membrane-bound proteins in eukaryotes—a glycan (oligosaccharide) is covalently attached to a nitrogen atom of the amino acid asparagine. Although it occurs as the protein is being translated, N-glycosylation is nevertheless classified as a post-translational modification [2]. N-glycosylation may influence protein folding and stability of the resulting glycoprotein [3].
|
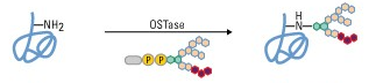
The process of N-glycosylation.
Oligosaccharide transferase (OSTase) mediates the transfer of a glycan molecule to a nitrogen atom in a protein. (Image source) |
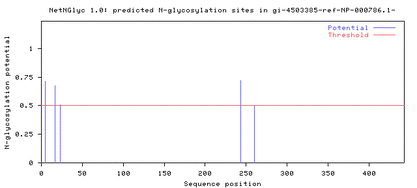
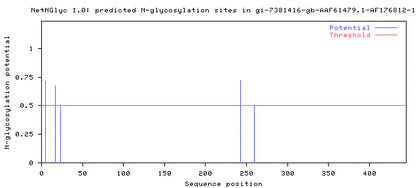
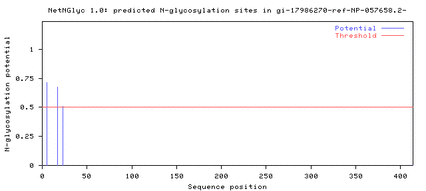
Comparing the protein sequences of the three DRD2 isoforms on NetNGlyc resulted in the following output, which shows that N-glycosylation sites are present near the 250th amino acid position in both the long isoform (D2L) and the longer isoform (D2(Longer)), but not in the short isoform (D2S) (Fig. 2).
Figure 1. N-glycosylation sites are present near the 250th amino acid position in D2L and D2(Longer).
Figure 2. The N-glycosylation sites near the 250th amino acid position
in D2L and D2(Longer) are absent in D2S.
in D2L and D2(Longer) are absent in D2S.
Discussion_________________________________________________________________________________________________
The data suggests that D2S may have altered function(s) compared to D2L and D2(Longer). Because its amino acid sequence is truncated relative to the other two isoforms, it is not surprising that glycosylation sites would be absent in D2S. D2S would therefore have a slightly altered protein structure relative to the longer isoforms. The data also supports findings from studies suggesting distinct physiological, signalling and pharmacological properties for different DRD2 isoforms [4].
The data suggests that D2S may have altered function(s) compared to D2L and D2(Longer). Because its amino acid sequence is truncated relative to the other two isoforms, it is not surprising that glycosylation sites would be absent in D2S. D2S would therefore have a slightly altered protein structure relative to the longer isoforms. The data also supports findings from studies suggesting distinct physiological, signalling and pharmacological properties for different DRD2 isoforms [4].
References
(1) Life Technologies. Protein Glycosylation. https://www.lifetechnologies.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/protein-glycosylation.html
(2) http://www.uniprot.org/help/carbohyd
(3) Imperiali B, O'Connor SE (1999). "Effect of N-linked glycosylation on glycopeptide and glycoprotein structure". Curr Opin Chem Biol 3 (6): 643–9.
(4) Beaulieu, J. M., & Gainetdinov, R. R. (2011). The physiology, signaling, and pharmacology of dopamine receptors. Pharmacological reviews, 63(1), 182-217.
(1) Life Technologies. Protein Glycosylation. https://www.lifetechnologies.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/protein-glycosylation.html
(2) http://www.uniprot.org/help/carbohyd
(3) Imperiali B, O'Connor SE (1999). "Effect of N-linked glycosylation on glycopeptide and glycoprotein structure". Curr Opin Chem Biol 3 (6): 643–9.
(4) Beaulieu, J. M., & Gainetdinov, R. R. (2011). The physiology, signaling, and pharmacology of dopamine receptors. Pharmacological reviews, 63(1), 182-217.